Computational Biology
Awesome Computational Biology  ¶
¶
A curated collection of databases, software, and papers related to computational biology.
Computational biology involves the development and application of data-analytical and theoretical methods, mathematical modelling and computational simulation techniques to the study of biological, ecological, behavioural, and social systems. — Wikipedia
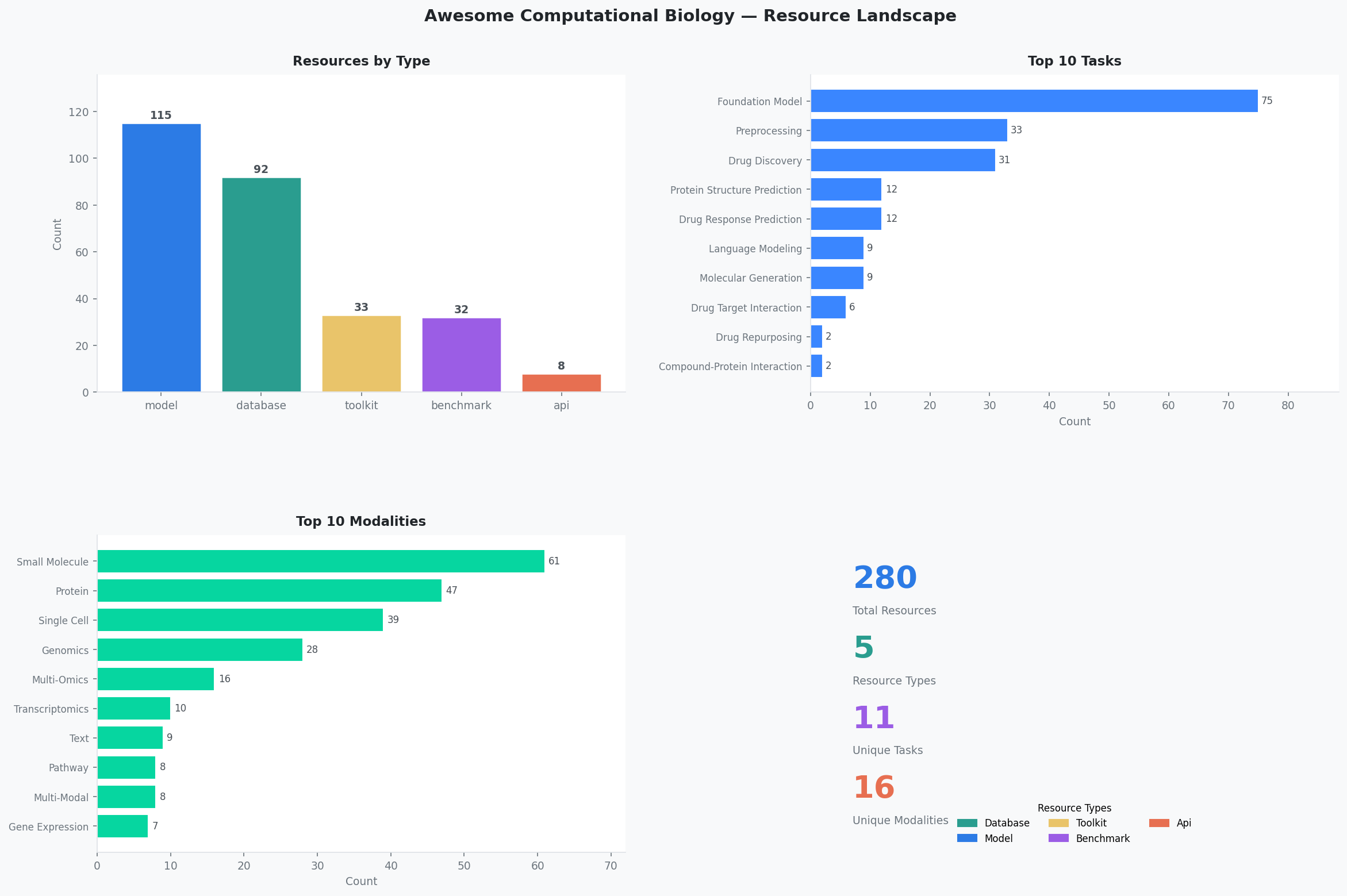
Overview¶
Interactive version: Resource Overview page
Regenerate the figure:python scripts/generate_overview.py
GitHub Pages UI¶
Browse and search the resources via the GitHub Pages UI.
- Search matches
name,description,tasks,modalities, andtags. - The Task, Modality, and Type filters map directly to
tasks,modalities, andtypeindocs/data/resources.json. - Clicking badges on cards applies the corresponding filter.
Databases¶
scRNA¶
- CZ CELLxGENE — Single-cell dataset repository and interactive explorer from the Chan Zuckerberg Initiative.
- Gene Expression Omnibus — Public functional genomics database.
- Human Cell Atlas — Open global atlas of all cells in the human body.
- Single Cell PORTAL — Public database for single-cell RNA.
- Single Cell Expression Atlas — Public database for single-cell RNA.
Compound¶
- PubChem — One of the largest chemical databases (compounds, genes, and proteins).
- ChEBI — Database focused on small chemical compounds.
- ChEMBL — Bioactive molecules with drug-like properties.
- ChemSpider — Chemical structure database.
- DrugTargetCommons — Community platform for curating and integrating experimental bioactivity data across drugs and targets.
- HMDB (Human Metabolome Database) — Comprehensive database of small molecule metabolites found in the human body.
- KEGG COMPOUND — Collection of small molecules and biopolymers.
- LIPID MAPS — Database of lipids.
- Rhea — Database of chemical reactions.
- DrugCentral — Online drug compendium with drug mode of action and indication information.
- Drug Repurposing Hub — Collections of drug repurposing data (drug, MoA, target, etc).
- Therapeutic Target Database — Drug-target, target-disease, and drug-disease datasets.
- ZINC ligand discovery database — Free database of commercially-available compounds for virtual screening.
Pathway¶
- PathwayCommons — Database of pathways and interactions.
- KEGG PATHWAY — Collection of pathway maps.
- WikiPathways — Database of biological pathways.
- Reactome — Expert-curated, peer-reviewed pathway database with detailed reaction mechanisms.
- BioCyc — Collection of pathway/genome databases across thousands of organisms.
- OmniPath — Comprehensive resource integrating protein interactions, signaling pathways, gene regulatory networks, and miRNA targets from over 100 databases.
- SIGNOR 2.0 — Database of causal signaling interactions and pathways, with signed and directed relationships between proteins.
- MSigDB (Molecular Signatures Database) — Curated gene sets derived from pathways and biological processes.
Mass Spectra¶
- MassBank — Open source databases and tools for mass spectrometry reference spectra.
- MoNA MassBank of North America — Meta-database of metabolite mass spectra, metadata, and associated compounds.
Protein¶
- THE HUMAN PROTEIN ATLAS — Comprehensive human protein database (cells, tissues, organs).
- PROTEIN DATA BANK (PDB) — 3D structures of proteins, nucleic acids, complexes.
- UniProt — Functional information on proteins.
- AlphaFold Protein Structure Database — 3D protein structure predictions.
- RCSB Protein Data Bank — Repository for structural data of biological molecules.
- Critical Assessment of Structure Prediction (CASP) — Assessing methods for protein structure prediction.
- Uniclust — Clustered protein sequence databases.
- UniRef — Non-redundant sequence database clustering UniProtKB entries at multiple sequence identity thresholds.
- CATH database — Hierarchical classification of protein domain structures.
- SAbDab — Structural Antibody Database containing all antibody structures in the PDB.
- OADB (Observed Antibody Space Database) — Database of antibody sequences from immune repertoire sequencing.
- InterPro — Protein families, domains, and functional sites database integrating 14 member databases including Pfam and PROSITE.
- Pfam — Database of protein families described by multiple sequence alignments and hidden Markov models.
- NeXtProt — Expert knowledge base on human proteins with deep functional annotation, complementary to UniProt.
Genome¶
- ENCODE — Encyclopedia of DNA Elements; regulatory and functional genomic elements across the genome.
- Ensembl — Genome browser and annotation database for vertebrate and other eukaryotic genomes.
- Human Genome Resources at NCBI — Database for genomics, proteomics, transcriptomics, and systems biology.
- GenBank — NCBI's database of genetic sequences.
- UCSC Genome Browser — UCSC's genome browser.
- cBioPortal — Cancer genomics database; aggregating many patient datasets.
- 10x Genomics Dataset — Collection of single-cell datasets.
- The Genotype-Tissue Expression (GTEx) — Human gene expression and regulation resource.
- Dependency Map (DepMap) — CRISPR-Cas9 screens in cancer cell lines.
- Catalogue Of Somatic Mutations In Cancer (COSMIC) — Resource on somatic mutations in cancers.
- MGnify — Resource for metagenomic and metatranscriptomic data.
- JASPAR — Database of transcription factor binding profiles.
- gnomAD — Genome Aggregation Database; genetic variation from large-scale sequencing projects.
- Rfam — Database of RNA families with sequence alignments and consensus structures.
- ROADMAP Epigenomics — Reference epigenome maps for 111 primary human cell types and tissues, including histone modifications, chromatin accessibility, and DNA methylation.
- FANTOM5 — Functional annotation of mammalian genome; comprehensive atlas of active enhancers, promoters, and transcription start sites across human and mouse cell types.
Disease¶
- KEGG DRUG — Comprehensive, approved drug information.
- DrugBank — Database of drugs and targets (University of Alberta).
- DisGeNET — Database of gene-disease associations integrating expert-curated and GWAS data.
- OMIM (Online Mendelian Inheritance in Man) — Comprehensive database of human genes and genetic disorders.
- Open Targets Platform — Systematic target identification and prioritization platform integrating genetics, genomics, and drug data for drug discovery.
- Human Phenotype Ontology (HPO) — Standardized vocabulary of phenotypic abnormalities in human disease, linking genes, variants, and clinical features.
- DISEASES — Gene–disease association database integrating evidence from text mining, curated databases, and experimental data.
Interaction¶
Drug-Gene Interaction¶
- DGIdb — Drug-gene interactions and the druggable genome.
- Comparative Toxicogenomics Database — Chemical-gene interactions, chemical-disease and gene-disease associations, chemical-phenotype associations.
- SNAP — Dataset of drug-gene interactions.
Drug (Cell Line) Response¶
- NCI60 — Focuses on 60 cancer cell lines and many drugs.
- Genomics of Drug Sensitivity in Cancer (GDSC) — Drug sensitivity for ~1000 human cancer cell lines and hundreds of compounds.
- Cancer Cell Line Encyclopedia — Database of ~1000 cancer cell lines.
- CellMiner Cross Database (CellMinerCDB) — Integrates multiple cancer cell line databases.
Chemical-Protein Interaction¶
- STITCH — Chemical-protein interactions.
- BindingDB — Compounds and target database.
- Davis kinase inhibitors DB — Experimental kinase inhibitor binding affinity dataset for protein–ligand interaction research.
- Kinase Inhibitor Bioactivity Data (KIBA) — Integrated bioactivity scores for kinase inhibitors combining Ki, Kd, and IC50 measurements.
- PDBBind — Binding affinity data for biomolecular complexes.
Protein-Protein Interaction¶
- STRING — PPI networks for multiple organisms.
- BioGRID — Protein, genetic, and chemical interactions.
- HIPPIE — Human protein-protein interaction database.
- IntAct — Open-source molecular interaction database and analysis system from EMBL-EBI.
Knowledge Graph¶
- Drug Mechanism Database (DrugMechDB) — Mechanisms of action from drug to disease.
- DRKG — Large-scale biological knowledge graph for drug discovery.
- Hetionet — Heterogeneous network integrating genes, diseases, drugs, pathways, and more.
- PrimeKG — Multi-modal precision medicine knowledge graph integrating clinical, genetic, and drug data.
Gene Regulatory Network¶
- TRRUST v2 — Manually curated database of human and mouse transcriptional regulatory interactions between transcription factors and their target genes, expanded with literature-derived evidence.
- RegNetwork — Database of gene regulatory networks covering transcription factor–target gene and miRNA–gene interaction data across multiple species.
- miRBase — Reference repository for microRNA gene annotations, sequences, and experimentally validated targets.
Clinical Trial¶
- ClinicalTrials.gov — Privately and publicly funded clinical studies.
- ICD10 — International Classification of Diseases, 10th revision.
- EU Drug Regulating Authorities Clinical Trials DB (EudraCT) — European clinical trial database.
- MIMIC-IV — Freely accessible critical care database.
Benchmarks & Datasets¶
- 1000 Genomes Project — Reference panel of human genetic variation from 2,504 individuals across 26 populations.
- BACE — Binary classification and regression dataset for β-secretase 1 (BACE-1) inhibitor binding affinity.
- BEAT AML — Functional ex vivo drug sensitivity measurements paired with genomics for acute myeloid leukemia.
- BindingDB Curated Sets — Curated binding affinity datasets for protein–ligand interaction benchmarking.
- Cancer Therapeutics Response Portal (CTRP) — Drug sensitivity profiles across ~900 cancer cell lines for >400 compounds.
- ClinTox — Clinical toxicity dataset contrasting FDA-approved drugs with those that failed clinical trials due to toxicity.
- CPTAC (Clinical Proteomic Tumor Analysis Consortium) — Multi-omic proteogenomic datasets for multiple cancer types linking proteomics with genomics.
- CrossDocked2020 — Large-scale dataset for structure-based virtual screening.
- DUD-E (Directory of Useful Decoys, Enhanced) — Structure-based virtual screening benchmark with active ligands and challenging decoy sets across diverse protein targets.
- FLIP (Fitness Landscape Inference for Proteins) — Benchmark collection of protein fitness landscape datasets for evaluating protein ML models.
- Genomics of Drug Sensitivity in Cancer (GDSC) — Drug sensitivity for ~1000 human cancer cell lines and hundreds of compounds.
- GuacaMol — Benchmark suite for generative molecular design models.
- JUMP Cell Painting Datasets — Consortium-scale cell imaging perturbation datasets (chemical and genetic) for phenotypic profiling and drug discovery research.
- LINCS L1000 — Gene expression profiles (978 landmark genes) for >20,000 chemical and genetic perturbations across cell lines.
- MoleculeNet — Benchmark datasets for molecular machine learning.
- MOSES — Benchmarking platform for molecular generation models.
- NCI60 — Drug sensitivity benchmark across 60 diverse human cancer cell lines.
- OGB (Open Graph Benchmark) — Large-scale graph ML benchmark suite including biological datasets such as ogbl-ppa (protein-protein associations) and ogbg-molhiv.
- OpenBioLink — Benchmark datasets for biological knowledge graph completion.
- PharmGKB — Curated pharmacogenomics dataset linking genetic variants to drug response phenotypes across thousands of drugs.
- PK-DB — Open database of experimental pharmacokinetics (PK) and ADME data from clinical and preclinical studies.
- PRISM — Cancer drug sensitivity profiling of >4,500 drugs across >900 cancer cell lines using pooled-cell-line barcoding.
- ProteinGym — Large-scale benchmark of deep mutational scanning assays for evaluating protein fitness landscape models.
- QM9 — Quantum chemistry properties for 134K stable small organic molecules computed at DFT level.
- scIB (Single-cell Integration Benchmarks) — Comprehensive benchmarking framework for single-cell data integration methods.
- scPerturb — Curated and continuously updated single-cell perturbation data resource spanning CRISPR and drug perturbation studies.
- SIDER (Side Effect Resource) — Database of 1,430 approved drugs with their recorded adverse drug reactions across 27 system-organ classes.
- Tabula Muris — Comprehensive single-cell atlas of 20 mouse organs and tissues, enabling cross-tissue and cross-species comparisons.
- Tabula Sapiens — Comprehensive human single-cell atlas of ~500K cells from 24 organs and tissues across multiple donors.
- TAPE (Tasks Assessing Protein Embeddings) — Benchmark suite of five biologically meaningful semi-supervised learning tasks for evaluating protein representations.
- The Cancer Genome Atlas (TCGA) — Comprehensive multi-omics (genomics, transcriptomics, proteomics, methylation) dataset for 33 cancer types across ~11,000 patients.
- Therapeutics Data Commons (TDC) — Unified benchmark suite covering ADMET, drug-target interaction, drug response, and more.
- Tox21 — 12,707 compounds tested in 12 nuclear receptor and stress-response pathway biochemical assays for toxicity prediction.
- UK Biobank — Large-scale biomedical database of ~500K participants with genetic, imaging, and health data for population genetics and disease studies.
API¶
- PubMed E-utilities (esearch/efetch) — APIs for searching and retrieving biomedical literature from PubMed.
- NCBI E-utilities — Unified APIs for accessing NCBI databases (Gene, GEO, SRA, PubChem, etc).
- UniProt REST API — Programmatic access to protein sequence and functional annotation data.
- Ensembl REST API — API for genomic annotations, variants, genes, and comparative genomics.
- KEGG REST API — API for accessing KEGG pathways, compounds, genes, and reactions.
- ChEMBL Web Services — REST API for bioactive molecules, targets, and bioassays.
- Open Targets Platform API — API for target–disease associations integrating genetics, genomics, and drug data.
- ClinicalTrials.gov API — API for querying clinical trial metadata and results.
Preprocessing Tools¶
- Chemistry Development Kit — Cheminformatics software & machine learning tools.
- Biopython — Collection of Python tools for biological computation including sequence analysis, structure parsing, and database access.
- FlashDeconv — High-performance spatial transcriptomics deconvolution (~1M spots in ~3 min).
- RDKit — Cheminformatics software & machine learning toolkit.
- DeepChem — Deep learning library for drug discovery, quantum chemistry, and materials science.
- ChatSpatial — MCP server for spatial transcriptomics analysis via natural language.
- Scanpy — Python library for scRNA-seq analysis.
- Seurat — R library for scRNA-seq analysis.
- scvi-tools — Probabilistic models for single-cell omics data analysis.
- CellTypist — Automated cell type annotation for scRNA-seq.
- Squidpy — Python library for spatial single-cell analysis.
- GROMACS — Molecular dynamics simulation package for biochemical molecules.
- MDAnalysis — Python library for analyzing and altering molecular dynamics simulation trajectories.
- OpenMM — High-performance toolkit for molecular simulation and GPU-accelerated MD.
- scVelo — RNA velocity estimation for single-cell transcriptomics, inferring the direction and speed of cell differentiation.
- STAR — Ultrafast universal RNA-seq aligner with support for spliced alignment and single-cell quantification via STARsolo.
- kallisto — Near-optimal RNA-seq quantification using pseudoalignment for fast transcript abundance estimation.
- Harmony — Fast and scalable integration of single-cell data across datasets, conditions, technologies, and species.
- Monocle3 — Single-cell trajectory analysis tool for learning developmental trajectories and ordering cells in pseudotime.
- CellChat — Inference and analysis of cell-cell communication ligand-receptor networks from single-cell transcriptomics data.
- SCENIC — Single-cell regulatory network inference and clustering linking transcription factors to co-expressed gene modules.
- DoubletFinder — Machine learning approach for detecting multiplet (doublet) artifacts in single-cell RNA-seq data.
- Numbat — Haplotype-aware copy number variation inference from single-cell RNA-seq using hidden Markov models.
- CaSpER — CNV identification and visualization by integrative analysis of single-cell or bulk RNA-seq data.
- CellCharter — Identification and characterization of spatial cell niches from spatial transcriptomics using VAEs and Gaussian mixture models.
- STAGATE — Adaptive graph attention auto-encoder for spatial domain identification in spatial transcriptomics.
- NCEM — GNN-based model for learning intercellular communication from spatial graphs of cells.
- DeepTalk — Graph attention network for deciphering cell-cell communication from spatial transcriptomics data.
- COMMOT — Optimal transport-based framework for screening cell-cell communication in spatial transcriptomics.
- TIGON — Neural optimal transport method for reconstructing growth and dynamic trajectories from single-cell transcriptomics.
- LINGER — Neural network for gene regulatory network inference from single-cell multiome (RNA+ATAC-seq) data with bulk data pretraining.
- sciPENN — RNN-based method for simultaneous protein expression prediction, uncertainty estimation, and cell-type label transfer from CITE-seq and scRNA-seq data.
- MOGONET — Multi-omics graph convolutional network framework for patient classification and biomarker identification.
Machine Learning Tasks and Models¶
Drug Discovery¶
Drug Response Prediction¶
- drGAT — Attention-based model for drug response prediction with gene explainability.
- MOFGCN — GCN + heterogeneous network.
- DeepDSC — Autoencoder + fully connected NN.
- DGDRP — Multi-view embedding neural network.
- DeepAEG — GNN embedding + attention mechanism.
- RECOVER — Machine learning framework for predicting synergistic drug combination responses across cell lines.
- TGSA — Tumor gene set and attention-based model leveraging biological pathway knowledge for drug response prediction.
- HiDRA — Hierarchical network model incorporating gene and pathway-level information for cancer drug response prediction.
- PRNet — Deep generative model for predicting transcriptional responses to novel chemical perturbations for drug discovery.
- chemCPA — Compositional perturbation autoencoder for predicting single-cell transcriptional responses to unseen drug perturbations and dose combinations.
- cycleCDR — Interpretable cycle-consistency framework for modeling cellular responses to drug perturbations.
- DRUML — Ensemble machine learning framework combining standard ML with deep learning to systematically rank anti-cancer drugs from proteomics and RNA-seq data.
Drug Repurposing¶
- DeepPurpose — Deep learning library for drug repurposing.
- TranSiGen — Dual-VAE architecture for ligand-based virtual screening, drug response prediction, and drug repurposing using chemical-induced transcriptional profiles.
Drug Target Interaction¶
- NeoDTI — Library for drug-target interaction prediction.
- DTINet — Network-based framework integrating heterogeneous biological data for DTI prediction.
- DeepDTA — Deep learning model using CNNs on protein sequences and drug SMILES.
- GraphDTA — Graph neural network–based DTI prediction using molecular graphs.
- MolTrans — Transformer-based DTI model leveraging molecular substructures.
- DrugBAN — Bilinear attention network for interpretable DTI prediction.
Compound-Protein Interaction¶
- MCPINN — Drug discovery via compound-protein interaction and machine learning.
- TransformerCPI — CPI prediction using Transformer.
Molecular Generation¶
- REINVENT — Reinforcement learning for de novo drug design.
- MolGPT — Transformer-based model for molecular generation.
- Molecular Transformer — Sequence-to-sequence model for retrosynthesis prediction.
- TargetDiff — 3D equivariant diffusion model for structure-based drug design.
- DiffDock — Diffusion generative model for molecular docking, predicting the binding pose of small molecules to protein targets.
- JTVAE — Junction tree variational autoencoder for molecular graph generation that guarantees chemical validity via a hierarchical tree decomposition.
- DiffSBDD — Equivariant diffusion model for structure-based drug design that generates molecules and binding conformations for protein targets.
- ReLeaSE — Deep reinforcement learning framework for de novo drug design combining a generative and predictive model.
- PaccMannRL — Reinforcement learning-based generative model for de novo hit-like anticancer molecule design from transcriptomic data.
LLM for Biology¶
- AI4Chem/ChemLLM-7B-Chat — LLM for chemical & molecular science.
- BioGPT — LLM for biomedical text generation.
- GeneGPT — LLM for biomedical information, integrated with various APIs.
- GenePT — Foundation LLM for single-cell data.
- scPRINT — Pretrained on 50M cells for scRNA-seq denoising & zero imputation.
- ClawBio — Bioinformatics-native AI agent skill library with local-first pharmacogenomics, ancestry PCA, semantic similarity, nutrigenomics, and metagenomics skills.
- BioMedLM — 2.7B parameter GPT-2-style language model trained exclusively on biomedical literature from PubMed for biomedical question answering and text generation.
- MolT5 — Language model for molecular tasks bridging text and SMILES, enabling molecule captioning and text-driven molecule generation.
- ChatDrug — LLM-based conversational pipeline for drug discovery, using natural language prompts for iterative drug editing and optimization.
Foundation Models¶
Single-cell Foundation Models¶
Transcriptomics Foundation Models¶
- scFoundation — Large-scale foundation model for single-cell gene expression, enabling multiple downstream tasks.
- scGPT — Transformer-based foundation model pretrained on millions of single-cell profiles.
- Geneformer — Context-aware, attention-based deep learning model pretrained on a large corpus of single-cell transcriptomes.
- BulkFormer — Foundation model for bulk RNA-seq data; learns general transcriptomic representations.
- scBERT — BERT-based foundation model pretrained on large-scale scRNA-seq data for cell type annotation.
- CellPLM — Cell pre-trained language model with inter-cell transformer architecture for diverse single-cell analysis tasks.
- UCE — Universal Cell Embeddings: zero-shot single-cell embedding model trained on 36M cells across species, tissues, and assays without fine-tuning.
- GEARS — Graph-based model for predicting transcriptional responses to single and combinatorial genetic perturbations using biological priors.
- SATURN — Transformer-based model integrating gene expression and protein sequences via a protein language model to learn unified multi-species cell embeddings.
- CancerFoundation — Single-cell RNA-seq foundation model trained exclusively on a curated dataset of malignant cells to learn cancer-specific embeddings.
Spatial Foundation Models¶
- GigaPath — Slide-level digital pathology foundation model pretrained on 1.3 billion pathology image tokens from whole-slide images.
- UNI — General-purpose self-supervised pathology foundation model trained on 100K+ whole-slide images for diverse computational pathology tasks.
- CONCH — Vision-language foundation model for computational pathology trained with contrastive captioning on pathology image–text pairs.
- Phikon — ViT-based pathology foundation model pretrained with iBOT self-supervision on TCGA whole-slide images.
- Nicheformer — Foundation model for single-cell and spatial omics using a transformer architecture with positional embeddings to encode spatial cell information.
- scGPT-spatial — Extension of scGPT for spatial transcriptomics with continual pretraining and a mixture-of-experts decoder for spatial gene expression analysis.
Multi-Omics Foundation Models¶
- scMulan — Single-cell multi-omic language model pretrained on ~10M cells spanning transcriptomics, epigenomics, and proteomics for cross-omics transfer tasks.
- totalVI — Probabilistic framework for joint analysis of paired scRNA-seq and protein (CITE-seq) data enabling multi-modal cell state representation across single-cell datasets.
- MultiVI — Multi-modal variational autoencoder for integrating paired and unpaired single-cell RNA-seq and ATAC-seq measurements into a unified latent space.
- MIRA — Probabilistic multimodal topic model jointly modeling single-cell transcriptomics and chromatin accessibility for regulatory network inference.
- GLUE — Graph-Linked Unified Embedding framework for unpaired single-cell multi-omics data integration across RNA, ATAC, methylation, and protein modalities.
- BABEL — Cross-modality translation model enabling prediction between scRNA-seq and scATAC-seq profiles without requiring paired single-cell measurements.
- Multigrate — Asymmetric multi-omics variational autoencoder for integrating single-cell data across RNA, ATAC, and protein modalities with missing-modality support.
- MOFA+ — Multi-Omics Factor Analysis framework identifying shared axes of variation across bulk and single-cell datasets including RNA, ATAC, proteomics, methylation, and copy number.
- GeneCompass — Large-scale foundation model integrating DNA regulatory sequences and single-cell transcriptomics from 120M+ cells across multiple species for gene regulation prediction.
- UnitedNet — Interpretable multi-task deep neural network for single-cell multi-omics integration spanning transcriptomics, chromatin accessibility, and proteomics.
- SpatialGlue — Graph attention network for spatial multi-omics integration jointly embedding spatial transcriptomics with chromatin accessibility or proteomics.
- MIDAS — Mosaic integration and differential accessibility model for single-cell multi-omics data that handles arbitrary missing-modality combinations across transcriptomics, chromatin accessibility, and proteomics.
- Concerto — Contrastive self-supervised learning framework for single-cell multimodal data integration, batch correction, and reference-query mapping.
- scButterfly — Dual-aligned variational autoencoder for single-cell cross-modality translation between paired and unpaired multiomics data.
- JAMIE — Joint variational autoencoder for multimodal single-cell data imputation and embedding.
- scPair — Bidirectional feedforward network for single-cell multimodal analysis with cross-modality prediction leveraging single-cell atlases.
Domain Alignment¶
- scArches — Transfer learning framework for mapping new single-cell datasets onto pre-trained reference atlases across batches, conditions, and modalities.
- TOSICA — Transformer-based framework for one-stop interpretable cell-type annotation supporting cross-dataset and cross-species transfer.
Compound Foundation Models¶
Compound Embedding¶
- ChemBERTa-2 — RoBERTa-based molecular language model pretrained on SMILES for small-molecule representation learning.
- GROVER — Self-supervised graph transformer for large-scale molecular representation learning from unlabeled compounds.
- Mol2Vec — Unsupervised molecular embedding method inspired by Word2Vec for learning vector representations of chemical substructures.
- MolFormer — Linear attention transformer pretrained on millions of SMILES strings for efficient molecular embeddings.
- Uni-Mol — 3D molecular pretraining framework for universal representation learning on molecules and protein pockets.
Protein Foundation Models¶
Pre-trained Embedding¶
- Evolutionary Scale Modeling (ESM) — Protein embeddings.
- ProtTrans — Suite of protein language models (ProtBERT, ProtT5, ProtXLNet) trained on billions of protein sequences from UniRef and BFD.
- ProGen2 — Protein language model trained on diverse protein families for sequence generation and fitness prediction.
- Ankh — Efficient protein language model optimized for downstream prediction tasks including secondary structure, localization, and function annotation.
Protein Structure Prediction and Design¶
- AlphaFold3 — Predicts structures of proteins, nucleic acids, small molecules, and their complexes.
- Boltz-1 — Open-source all-atom biomolecular structure prediction model for proteins, nucleic acids, small molecules, and their complexes achieving AlphaFold3-level accuracy.
- Chai-1 — Unified molecular structure prediction model covering proteins, nucleic acids, small molecules, and complexes.
- ESM3 — Multimodal protein language model that jointly reasons over sequence, structure, and function for generative protein design and engineering.
- ESMFold — Fast protein structure prediction using language model embeddings.
- RFdiffusion — Generative model for protein backbone design using diffusion.
- ProteinMPNN — Deep learning model for protein sequence design given backbone structure.
- OmegaFold — High-resolution de novo protein structure prediction from sequence.
- RoseTTAFold — Three-track neural network for protein structure prediction.
- OpenFold — Trainable, memory-efficient open-source reproduction of AlphaFold2 enabling custom protein structure prediction workflows.
- SaProt — Structure-aware protein language model using structure-aware tokens that encode both sequence and backbone geometry for improved function prediction.
- EvoDiff — Discrete diffusion framework for protein sequence generation trained on evolutionary-scale data, supporting unconditional generation, disordered region design, and functional motif scaffolding. [ paper-2023 ]
Multi-Modal Foundation Models¶
- CHIEF — Clinical Histopathology Imaging Evaluation Foundation model integrating histology images and clinical context for pan-cancer analysis.
- BiomedCLIP — CLIP-based vision-language foundation model for biomedical images and text trained on PubMed figure–caption pairs.
- PORPOISE — Pan-cancer integrative histology-genomic analysis framework using multimodal deep learning for patient stratification.
- PathomicFusion — Integrated framework fusing histopathology and genomic features via CNN, GNN, and attention gating for cancer diagnosis and prognosis.
- Virchow — Million-slide digital pathology foundation model using a vision transformer and self-supervised distillation for tile-level pathology image representation.
- TOAD — Tumor Origin Assessment via Deep-learning; weakly-supervised multi-task model predicting cancer primary origin from H&E whole-slide images.
- PLIP — Vision-language foundation model for pathology trained with contrastive learning on pathology image–text pairs for image classification and text-to-image retrieval.
- MUSK — Vision-language foundation model for precision oncology analyzing multimodal paired text and pathology image data for biomarker prediction and retrieval.
Genomics Foundation Models¶
- Nucleotide Transformer — Foundation model for genomic sequences across multiple species.
- DNABERT — Pre-trained bidirectional encoder for DNA sequence analysis.
- DNABERT-2 — Improved genome foundation model with efficient tokenization.
- Enformer — Transformer model predicting gene expression from DNA sequence.
- Basenji — Sequential regulatory activity prediction from DNA sequences.
- Caduceus — Bidirectional equivariant long-range DNA sequence model based on Mamba.
- Evo — Long-context genomic foundation model (up to 1M tokens).
- HyenaDNA — Long-range genomic foundation model handling sequences up to 1M tokens with sub-quadratic attention.
- Borzoi — Extended successor to Enformer for predicting RNA-seq coverage from long genomic sequence windows (524 kb) with improved resolution.
- DeepSEA — Deep learning framework for predicting chromatin effects of sequence alterations with single-nucleotide sensitivity across thousands of chromatin features.
- Sei — Sequence-to-function framework learning a genome-wide regulatory activity code from DNA sequences for variant effect prediction.
- GPN (Genomic Pre-trained Network) — Masked language model for DNA sequences enabling zero-shot variant effect prediction without requiring functional annotations.
Citation¶
If you use this list in papers, slides, or documentation, please cite this repository via CITATION.cff (also available through GitHub's Cite this repository button).
Curation Criteria (Strict)¶
To keep quality high, additions should meet all of the following:
- The resource is trustworthy and relevant to computational biology.
- The primary link points to an official source (official docs, organization site, maintained repository, or official dataset page).
- The resource has evidence of technical substance: ideally a peer-reviewed paper; at minimum a preprint or official technical documentation.
- The description is factual and concise (no marketing copy).
- Duplicate or near-duplicate entries should be avoided.
We generally do not accept entries that are only promotional pages, personal opinion posts, or generic blog posts without technical references.
Update & Link Rot Policy¶
- Link validity is monitored by the Link Check workflow.
- If a link repeatedly fails, maintainers may replace it with an official mirror/canonical URL or remove the entry until a stable URL is available.
- Contributions fixing broken links are welcome and encouraged.
Data Schema & Contribution Workflow¶
- Data schema reference:
docs/data/SCHEMA.md. - Source-of-truth workflow:
- Edit/add resources in
README.md. - Regenerate machine-readable artifacts:
python scripts/sync_resources_from_readme.pypython scripts/build_resources.py
- Commit updated data files (
data/resources.yml,data/resources.json,data/resources.csv,docs/data/resources.json) with your README change. - Contribution guide:
contributing.md.